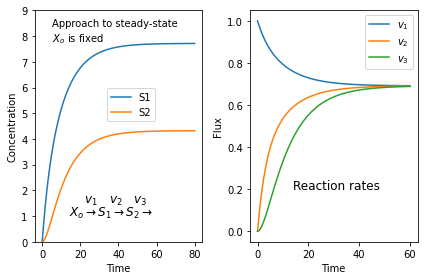
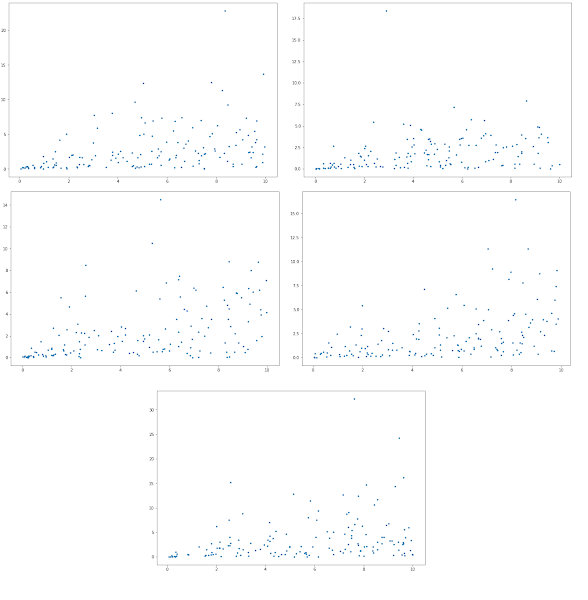
Here is a simple script that will generate a large number of random mass-action models and plot simulations of each in a grid.
Each grid has a number written on it so that you can grab the associated model if you find some interesting beahvior. In this case we see model 81 is oscillatory. You'll need the teUtils package for this in order to access the random network generator.
import tellurium as te
import roadrunner
import teUtils as tu
import matplotlib.pyplot as plt
import numpy as np
numRows = 15
numCols = 15
plt.subplots(numRows, numCols, figsize=(19,16))
count = 0
models = []
while count < numRows*numCols:
model = tu.buildNetworks.getRandomNetwork(10,20)
r = te.loada(model)
try:
m = r.simulate (0, 160, 200)
try:
models.append (model)
if count % 20 == 0:
print (count)
ax = plt.subplot (numRows, numCols, count+1)
ax.tick_params(axis='both', which='major', labelsize=7)
ax.tick_params(axis='both', which='minor', labelsize=7)
ax.set_xlabel('Time')
te.plotArray(m, show=False)
maxy = ax.get_ylim ()
ax.text (50, maxy[1]/2, str(count), fontsize=14)
count = count + 1
except:
# failed to find a steady state so probably a bad model
pass
except:
print ('Something very wrong with the model')
plt.show()
There is one run I did and you'll model 81 ihas some intersting dynamics:
This is model 81 pulled out and resimulated to show the dynamics more clearly:
import tellurium as te
# Get the model using:
# print (models[81])
# Then copy and paste the model as below:
r = te.loada("""
var S0, S1, S2, S4, S5, S6, S7, S8, S9
ext S3;
J0: S9 + S8 -> S6; E0*(k0*S9*S8);
J1: S6 -> S7 + S9; E1*(k1*S6);
J2: S2 + S0 -> S8; E2*(k2*S2*S0);
J3: S4 + S4 -> S6; E3*(k3*S4*S4);
J4: S5 -> S2 + S1; E4*(k4*S5);
J5: S9 -> S4 + S0; E5*(k5*S9);
J6: S1 -> S0 + S5; E6*(k6*S1);
J7: S0 -> S4 + S4; E7*(k7*S0);
J8: S3 -> S9; E8*(k8*S3);
J9: S5 -> S1 + S9; E9*(k9*S5);
J10: S5 -> S0 + S4; E10*(k10*S5);
J11: S5 -> S4 + S9; E11*(k11*S5);
J12: S5 + S1 -> S0; E12*(k12*S5*S1);
J13: S2 + S9 -> S4; E13*(k13*S2*S9);
J14: S1 -> S8; E14*(k14*S1);
J15: S8 + S0 -> S1; E15*(k15*S8*S0);
J16: S7 -> S8; E16*(k16*S7);
J17: S8 + S6 -> S7; E17*(k17*S8*S6);
J18: S4 -> S6 + S6; E18*(k18*S4);
J19: S5 -> S1 + S4; E19*(k19*S5);
k0 = 0.6158; k1 = 0.0524; k2 = 0.7206; k3 = 0.0261
k4 = 0.4946; k5 = 0.2428; k6 = 0.3249; k7 = 0.4854
k8 = 0.6743; k9 = 0.6320; k10 = 0.8954; k11 = 0.435
k12 = 0.580; k13 = 0.0298; k14 = 0.850; k15 = 0.342
k16 = 0.556; k17 = 0.3221; k18 = 0.584; k19 = 0.681
E0 = 1; E1 = 1; E2 = 1; E3 = 1
E4 = 1; E5 = 1; E6 = 1; E7 = 1
E8 = 1; E9 = 1; E10 = 1; E11 = 1
E12 = 1; E13 = 1; E14 = 1; E15 = 1
E16 = 1; E17 = 1; E18 = 1; E19 = 1
S3 = 1
S0 = 6; S1 = 3; S2 = 3; S4 = 2
S5 = 6; S6 = 2; S7 = 6; S8 = 2
S9 = 1
""")
m = r.simulate (0, 180, 100)
r.plot()
Here is the simulation from running the above code. Note that not every reaction is contributing to this behavior. You can remove J2, J3, J4, J6, J9, J10, J11, J12, J13, J14, J15, and J19 and the system will still oscillate.
What you can now is use another function in teUtils to look at a single model and plot a grid of simulations using random parameter values for that model. To do this just call plotRandSimGrid as follows. We apply it to model 81:
r = te.loada (models[81])
tu.plotting.plotRandSimGrid (r, endTime-500, ngrid=8, maxRange=1)
maxRange limits the range of random parmateer values. In this model, if the range is bigger, sometimes we get bad parameter sets which don't simulate. A maxRange of 1 seems to gives simulatable models. Most of the time the defaults are sufficient and you only have to pass in the roadrunner object, r.
Here is a sample run: